Virtual human modeling for modern clinical development
Iterate faster, test more broadly and expand what’s possible in trial design and beyond.
Our models guide smarter decisions
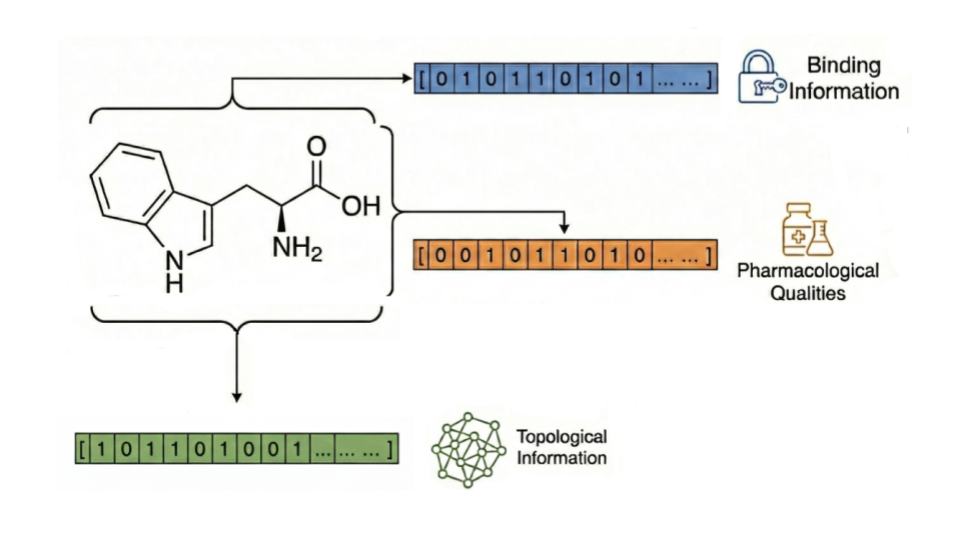
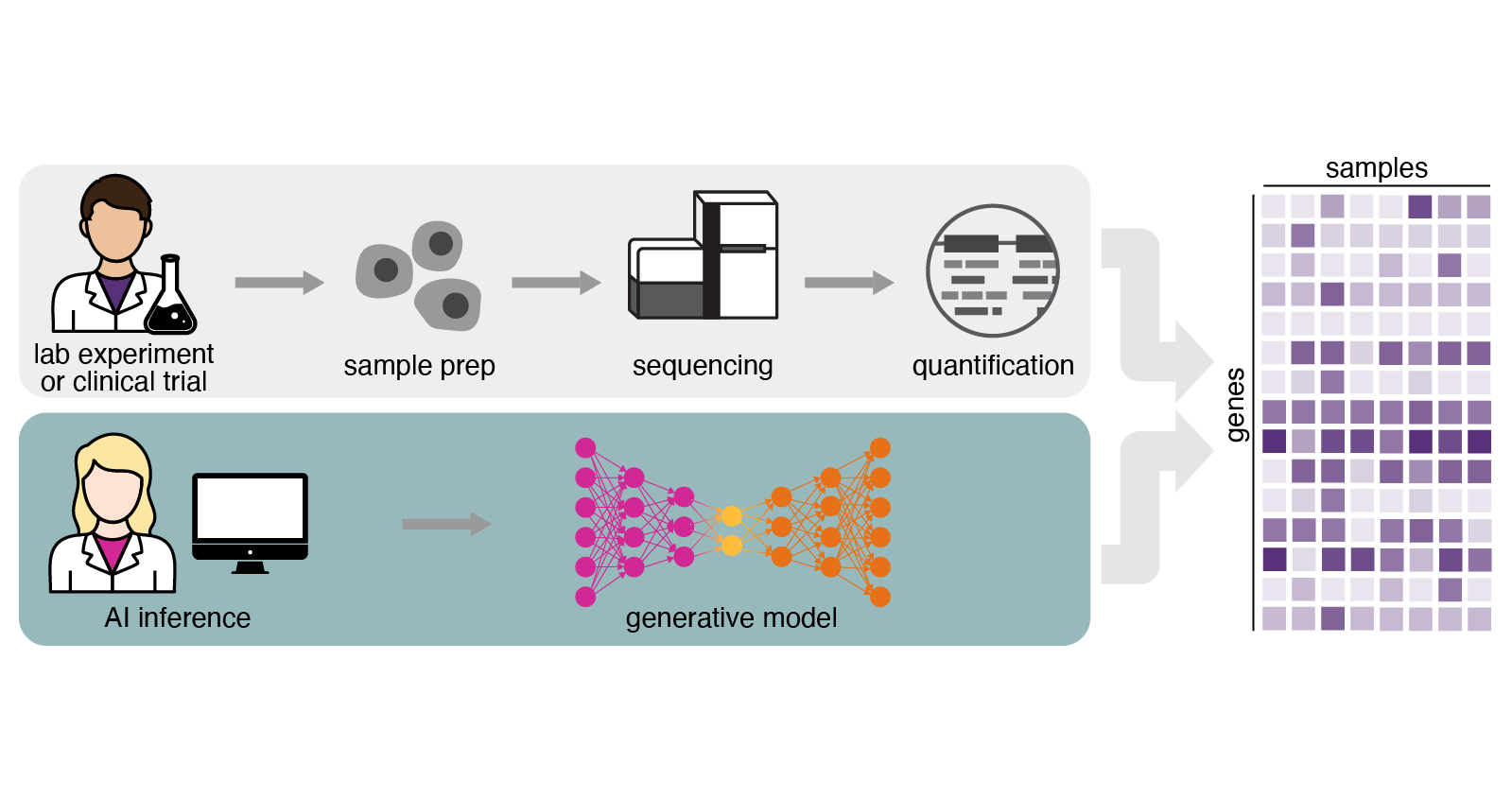
Our generative genomics models extend what scientists can examine by predicting how human tissues and cells respond to drugs, diseases and perturbations.
Describe experiments in natural language, and our model produces lab-quality data with results grounded in the largest annotated RNA-seq corpus ever assembled.
Unlock actionable clinical predictions
Small Phase 1/2 datasets leave critical questions unanswered. Our models extend your cohorts with synthetic patients matched to your trial profiles, enabling you to test endpoints, refine stratification, and model dose-response before committing hundreds of millions to pivotal studies.
Test trial designs in silico before protocol lock. Model patient responses across enrollment criteria and genetic backgrounds to refine inclusion/exclusion criteria, identify subgroups most likely to benefit, and explore endpoint strategies, de-risking pivotal trials early.
Rescue insights from negative trials by augmenting limited datasets. Identify responder subgroups, generate hypotheses for follow-up studies, or build evidence for alternative indications, turning failures into strategic pivots.
Predict real human tissue response
Identify off-target effects and tissue toxicities that preclinical models miss. Predict transcriptional responses across human tissue types to anticipate adverse events, prioritize safety biomarkers, and design monitoring strategies, catching signals before they threaten your program.
Understand how your therapeutic behaves in human tissues before committing to expensive studies. Focus resources on paths with the strongest biological rationale and clinical potential.
Map gene expression profiles across species to validate animal findings and increase translational success, reducing the risk of preclinical candidates failing in human trials.
Transform small datasets into development intelligence
Turn underpowered rare disease datasets into actionable intelligence. Generate synthetic cohorts that expand sample sizes while preserving biological characteristics of limited patient populations.
Identify and validate predictive biomarkers before large clinical studies. Generate gene expression profiles across patient subpopulations and disease states, enabling you to test stratification hypotheses, refine companion diagnostics, and design enrichment trials with confidence.
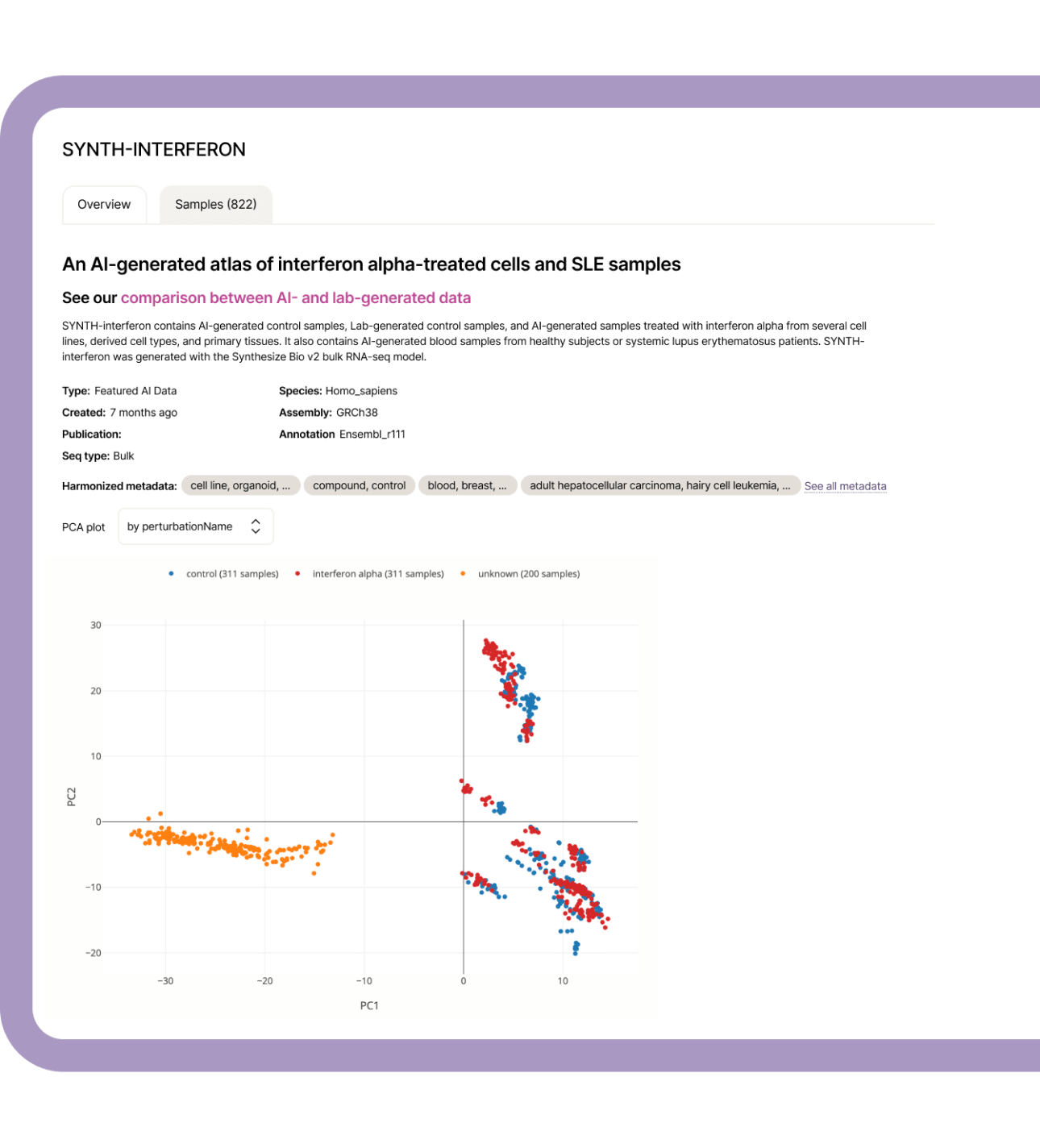
Collaborate across organizations using synthetic cohorts that mirror real patient data without patient-identifying information. Explore our SYNTH-cancer database—over 10,000 AI-simulated samples spanning 27 cancer types.

Foundation models designed for biopharma
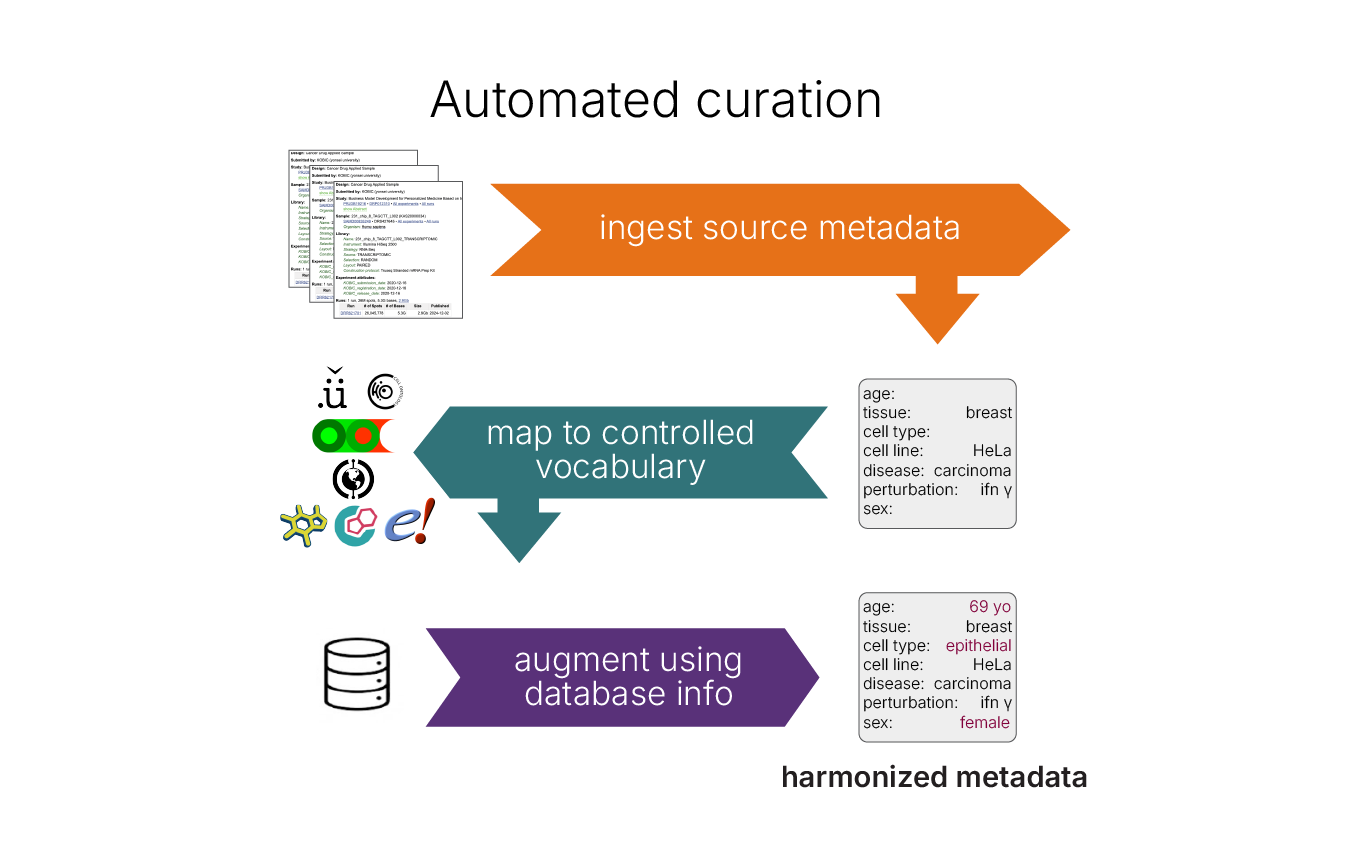
Read our preprint to dig deeper on the core GEM-1 model architecture, understand our rigorous validation standards and learn how we harmonized public data resources to create the largest annotated RNA-seq corpus ever assembled.
Let's accelerate your programs together
Partner with us
Work with us to apply our generative genomics capabilities to your therapeutic pipeline and trial design.


Experience GEM models
Try the model
Our public model accurately simulates a wide range of experiments based on your experimental description. For clinical applications and proprietary datasets, partner with us.


Your questions answered
GEM-1 is a foundation model that predicts human tissue responses to therapeutic interventions by generating lab-quality bulk and single-cell RNA-seq data from natural-language experimental descriptions. It's built on the largest well-annotated and consistently normalized RNA-seq corpus ever assembled, enabling predictions grounded in real human biology.
Drug development teams looking to de-risk clinical decisions, accelerate timelines, and make better-informed development choices:
Clinical Development Teams: De-risk trial design, identify predictive biomarkers, stratify patients by response likelihood, and extend early-phase data to inform pivotal trial decisions.
Translational Medicine: Validate mechanism of action in human tissues, predict safety signals before trials, and bridge preclinical findings to human biology.
Rare Disease Programs: Overcome small sample size limitations by augmenting limited patient datasets with biologically grounded synthetic cohorts.
Portfolio Strategy Teams: Extract value from failed trials, evaluate alternative indications, and build evidence for label expansions without waiting years for real-world data.
We partner with biotech and pharmaceutical companies to integrate gene expression predictions into clinical development programs. Partnerships are tailored to your specific needs, whether that's a focused project addressing a particular development challenge or ongoing collaboration across your pipeline. Partnership capabilities include custom model development, integration into your workflows, and dedicated scientific support. Your proprietary data remains private and is not used to train our models. Contact us to discuss how GEM models could support your programs.
GEM-1's prediction accuracy meets or exceeds the reproducibility standards of experimental biology. Our predictions correlate with real experimental outcomes with higher accuracy than the consistency of biological replicates measured across different labs.
Development teams use GEM-1 to design better trials: selecting enrollment criteria that enrich for responders, identifying predictive biomarkers for stratification, optimizing dose selection across patient subgroups, and anticipating safety signals in tissues that can't be routinely biopsied. The result: fewer late-stage trial failures and faster paths to approval by making data-driven decisions when real patient data is limited, expensive, or impossible to obtain.
GEM-1 data and predictions are designed to inform internal development decisions. Insights from GEM-1 can help you design stronger trials, identify better biomarkers, and build more compelling regulatory packages by optimizing trial design and patient selection strategies before committing to expensive studies. Currently, results from GEM-1 cannot serve as primary evidence in regulatory submissions.
Meet our team


Robert Bradley, PhD


Jeff Leek, PhD


Alice Walsh, PhD


Gregory Koytiger, PhD


Ariel Brumbaugh, PhD


Stein Setvik


Liza Ray, MBA





















News from Synthesize
Join us at the intersection of computation and biology
We’re building an inclusive team that’s inspired to transform scientific processes and human health.